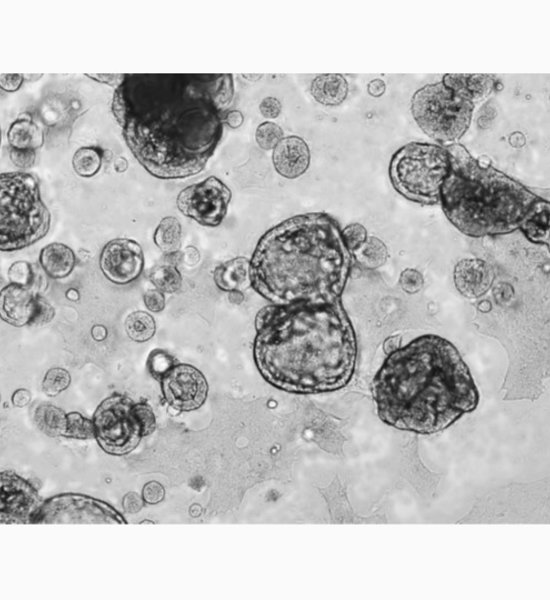
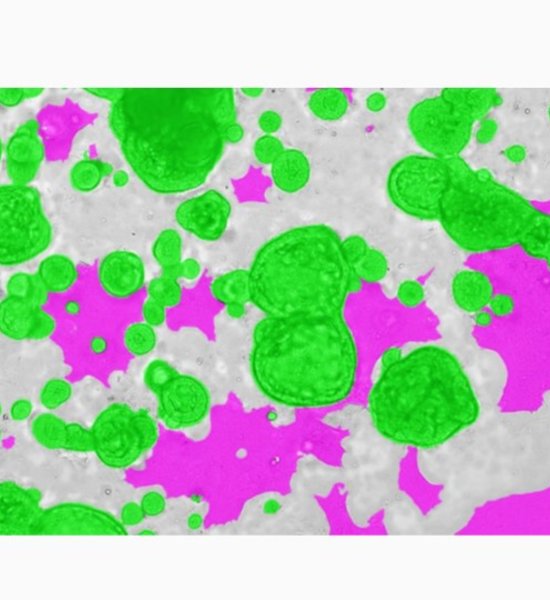
In recent years, research on organoids in regenerative medicine and drug discovery research has developed rapidly. In particular, the research for producing intestinal organoids from induced pluripotent stem cells (iPS cells) and human biological tissues has been remarkably developed, and the culture conditions for each have been almost established. However, imaging and image analysis methods of organoids have not been established yet. Organoids are composed of heterogeneous cell populations, and in addition to exhibiting a non-uniform morphology for each organoid, cell populations other than the evaluation target are likely to be included in the image. For these reasons, only by using image processing with a simple parameter threshold, high-precision segmentation of organoids cannot be performed, and quantification by image analysis is difficult. In the case of image analysis using a fluorescent label, there is a possibility that the above problems can be overcome, but today, the importance of label-free analysis without using a fluorescent label is increasing. In general, a three-dimensional culture method with an extracellular matrix is used for organoid culture, which makes imaging itself and image analysis more difficult. Therefore, in this study, we attempted a bright field image analysis of human iPSC-derived intestinal organoid using deep learning technology, which utilized in various fields. Since individual organoids cultured in Matrigel are located at different heights, all-in-focus images were obtained by Z-stack imaging. By using deep learning on the acquired image, only organoids exhibiting specific morphologies were segmented from the image, and feature quantities such as organoid count, diameter and area were calculated. Cell3iMager duos (SCREEN Holdings Co., Ltd.) was used for this imaging and image analysis. In this presentation, we will discuss the label-free analysis of organoids and their applications.